Proteomics
Proteomics Research
The Proteomics Facility at the University of Bradford is equipped with the latest instrumentation and provides research support in 3 main areas:
- identification of new protein targets for drug development and new biomarkers for disease detection and interpretation,
- functional and biochemical characterisation of established targets to assist refinement of next generation drug candidates,
- pharmacoproteomics and toxicoproteomics of drugs tested in cell lines, preclinical models and Phase I clinical trials, to determine their effects on protein expression, in liver, tumours and other organs.
Please contact us if you wish to discuss your proteomics needs or collaborative research opportunities with our team.
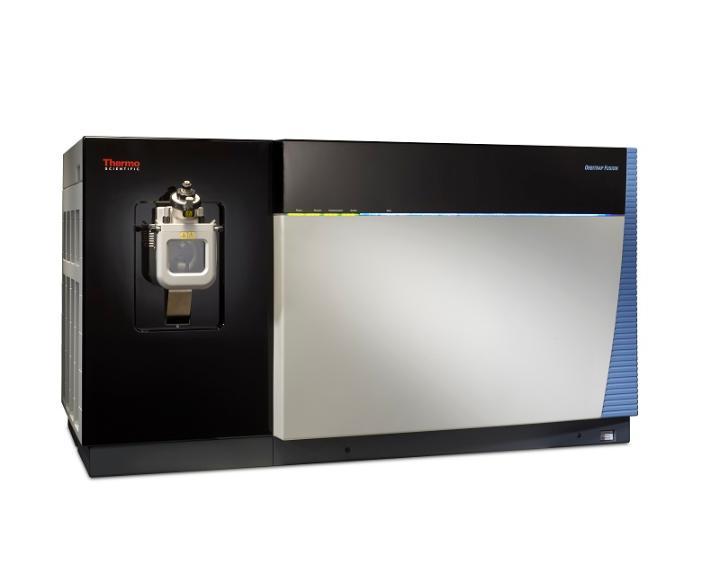

Thermo Scientific Orbitrap Fusion with Easy-Max NG source

-457x363.png)
-457x272.png)

-457x177.jpg)
